HDC PROTEIN PHYLOGENY
PROTEIN PHYLOGENY
Biochemical makeup, gene sequence, and function data imply that all organisms are genetically related. Phylogeny refers to the history of organisms and when their structures begin to diverge from one another. Specifically protein phylogeny is the history of a protein that has changed to suit a different organism. Referring back to homology, the protein sequences in each organism are not identical and therefore may not function identically either. Phylogeny helps us illustrate the changes in a protein in relation to the history of organisms [6].
HDC PROTEIN PHYLOGENY
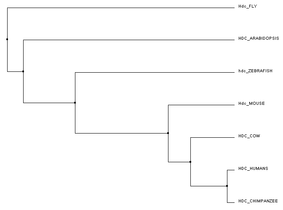
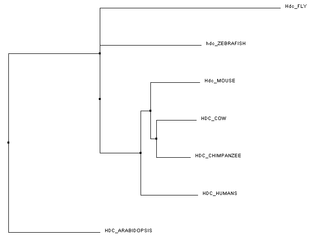
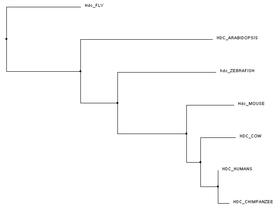
Below are trees created using the Clustal Omega program [3]. For the trees below, each species specific FASTA protein sequence was input and sent for alignment. Using the FASTA sequences allowed for fast matching. A FASTA sequence is a text-based format for representing, in this case, protein sequences, where amino acids are represented by single letter codes [2]. Each tree was formed using different comparison mechanisms. Figure 1 was created using the average distance between their protein sequences. Closely related sequences are on the same node and branch length is calculated from the distances between sequences. PID (percentage identity) is the similarity between each pair of sequences [4]. Figure 2 and Figure 3 both were created using the neighbour joining tree calculation. Neighbour joining tree form is an algorithm to find the tree with the shortest branch lengths. Figure 2 uses BLOSUM62 to measure distance instead of PID. BLOSUM62 (Blocks Substitution Matrix) gives higher scores when similar amino acids are substituted. These sequences are not identical but still contain some uniformity [5].
ANALYSIS
I believe the PID default platform is the easiest to view. It uses the percent identity of histidine decarboxylase in determining the claves on the trees. Mammals are grouped together followed by Arabidopsis and the fruit fly. BLOSUM62 shows us the similarities between amino acids. Because the amino acid length of Hdc in the fruit fly is much longer than the other organisms, the clade in the BLOSUM62 tree is farther from mammals than the PID platform trees.
Header Image Credit http://academic.reed.edu/biology/professors/srenn/pages/teaching/web_2010/MDCM_dinoflagellates/phylogeny.html
REFERENCES
1 "Dictionary" Merriam-Webster. Retrieved 13 Feb 2014 http://www.merriam-webster.com/dictionary/phylogeny
2 "FASTA format" Wikipedia: The Free Encyclopedia. Retrieved 13 Feb 2014 http://en.wikipedia.org/wiki/FASTA_format
3 "Clustal Omega" EMBL-EBI. Retrieved 13 Feb 2014 https://www.ebi.ac.uk/Tools/msa/clustalo/
4 "Calculation of trees from alignments" Jalview Help. Retrieved 13 Feb 2014 http://www.jalview.org/help/html/calculations/tree.html
5 "BLOSUM62 Miscalculations improve search performance" Styczynski et al. 2008 Nature Biotechnology Issue 26, p274-75. Retrieved 19 Feb 2014 http://www.nature.com/nbt/journal/v26/n3/full/nbt0308-274.html 6 "What is Phylogeny?" Tree of Life web project. Retrieved 19 Feb 2014 http://tolweb.org/tree/learn/concepts/whatisphylogeny.html
REFERENCES
1 "Dictionary" Merriam-Webster. Retrieved 13 Feb 2014 http://www.merriam-webster.com/dictionary/phylogeny
2 "FASTA format" Wikipedia: The Free Encyclopedia. Retrieved 13 Feb 2014 http://en.wikipedia.org/wiki/FASTA_format
3 "Clustal Omega" EMBL-EBI. Retrieved 13 Feb 2014 https://www.ebi.ac.uk/Tools/msa/clustalo/
4 "Calculation of trees from alignments" Jalview Help. Retrieved 13 Feb 2014 http://www.jalview.org/help/html/calculations/tree.html
5 "BLOSUM62 Miscalculations improve search performance" Styczynski et al. 2008 Nature Biotechnology Issue 26, p274-75. Retrieved 19 Feb 2014 http://www.nature.com/nbt/journal/v26/n3/full/nbt0308-274.html 6 "What is Phylogeny?" Tree of Life web project. Retrieved 19 Feb 2014 http://tolweb.org/tree/learn/concepts/whatisphylogeny.html
|
University of Wisconsin – Madison
Spring 2014 Genetics 564 |
|